-
Notifications
You must be signed in to change notification settings - Fork 2
Description
I'm solving EDMFT with CTINT as the impurity solver. The self-consistency for this requires not only the standard hybridization self-consistency, but also that the dynamical spin-spin interaction is consistent with the dynamical spin-spin correlator. So I'm using CTINT with both the dynamical spin-spin (Jperp) and the density-density (D0) turned on.
I'm finding that in the parameter region where CTINT has sign problems, it also does not seem to handle the spin asymmetry as expected (the alpha tensor, set by the solve parameter delta). A short EDMFT script that reproduces this issue for beta=30, J=0.5, U=6.0 is attached.
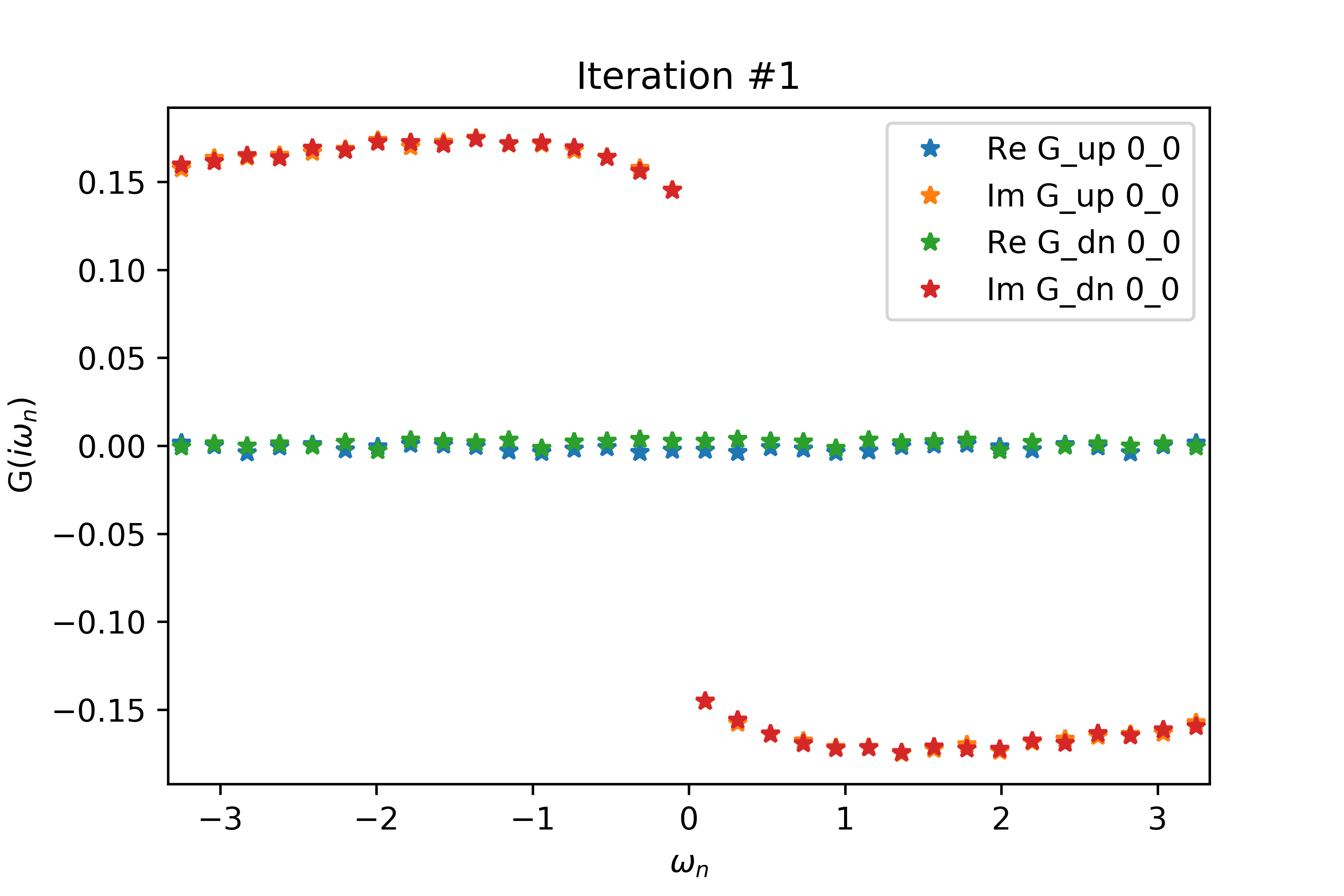
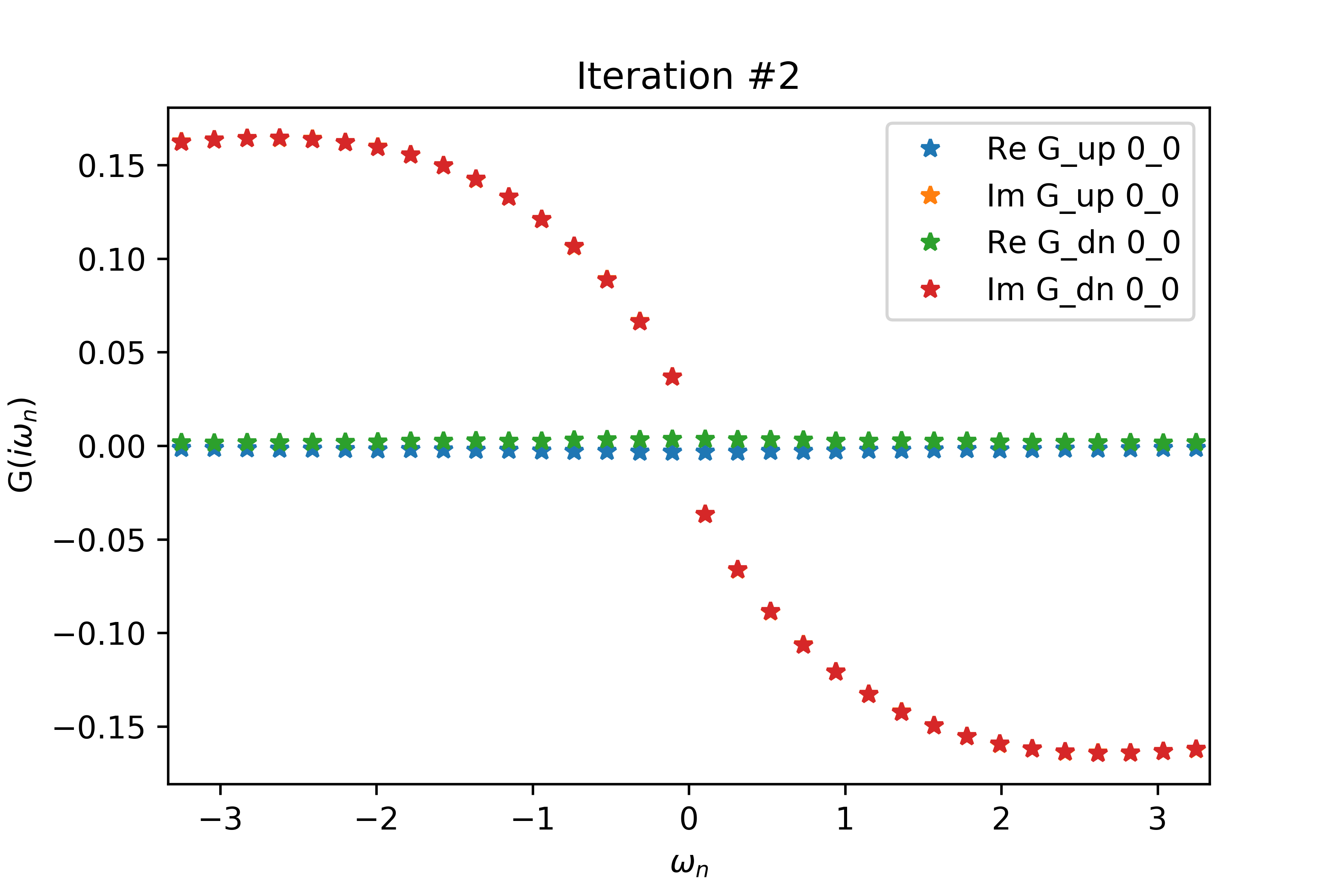
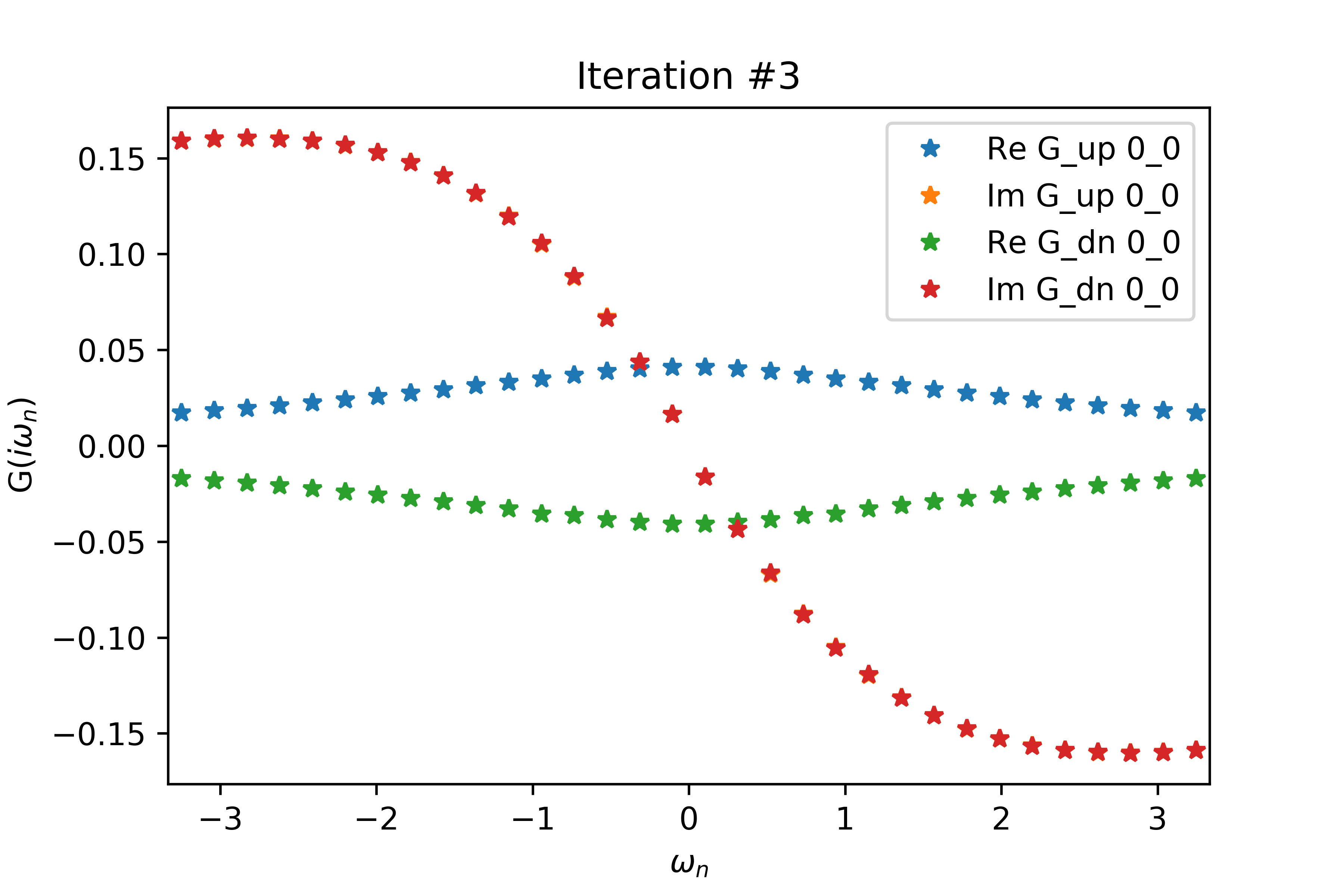
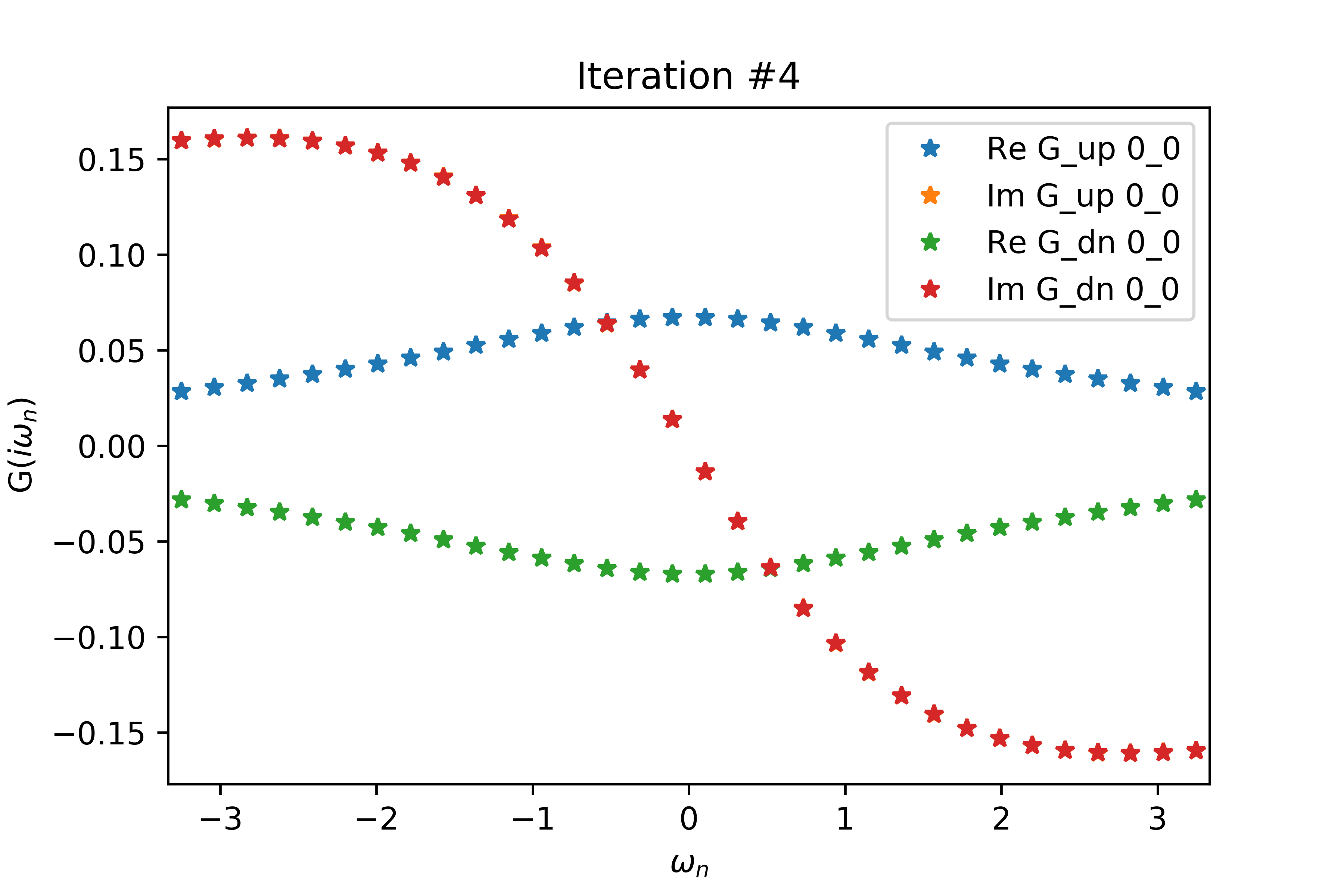
Even for a relatively small value of delta (delta=.2 below), the spin-up and spin-down interacting Green's functions have non-zero real parts related by a negative sign, when both G's should have vanishing real part. Increasing delta also increases the magnitudes of the non-zero real parts.
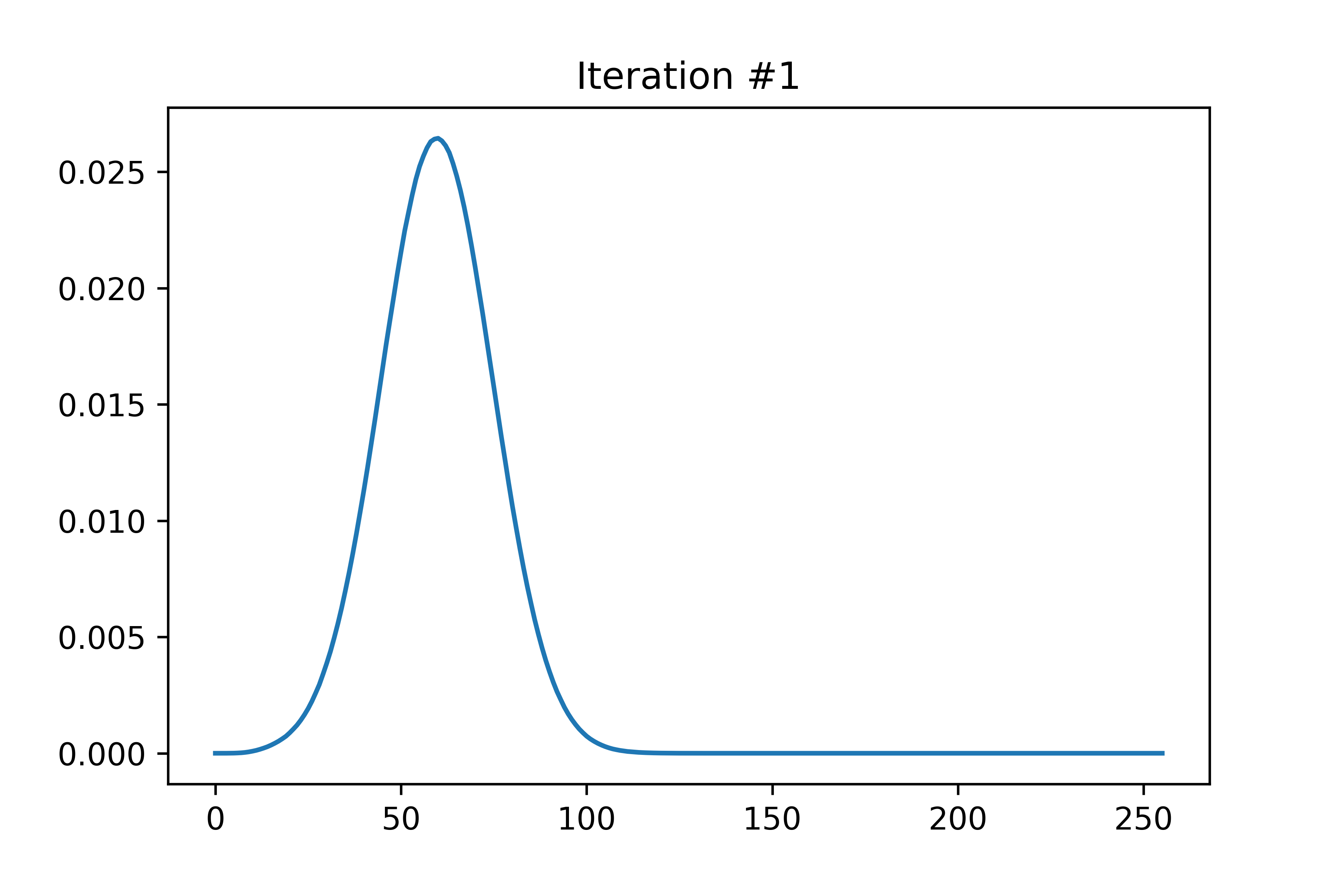
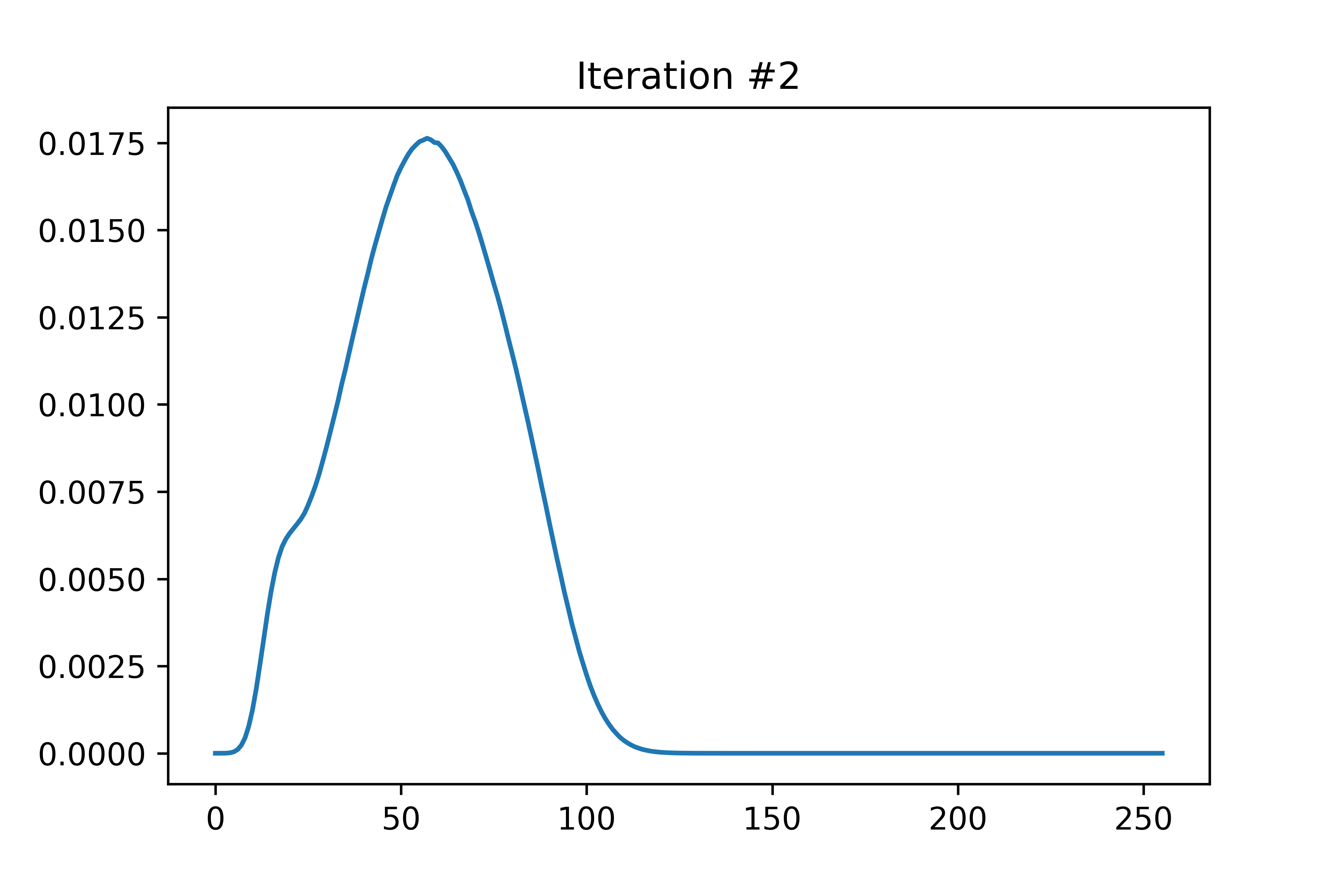
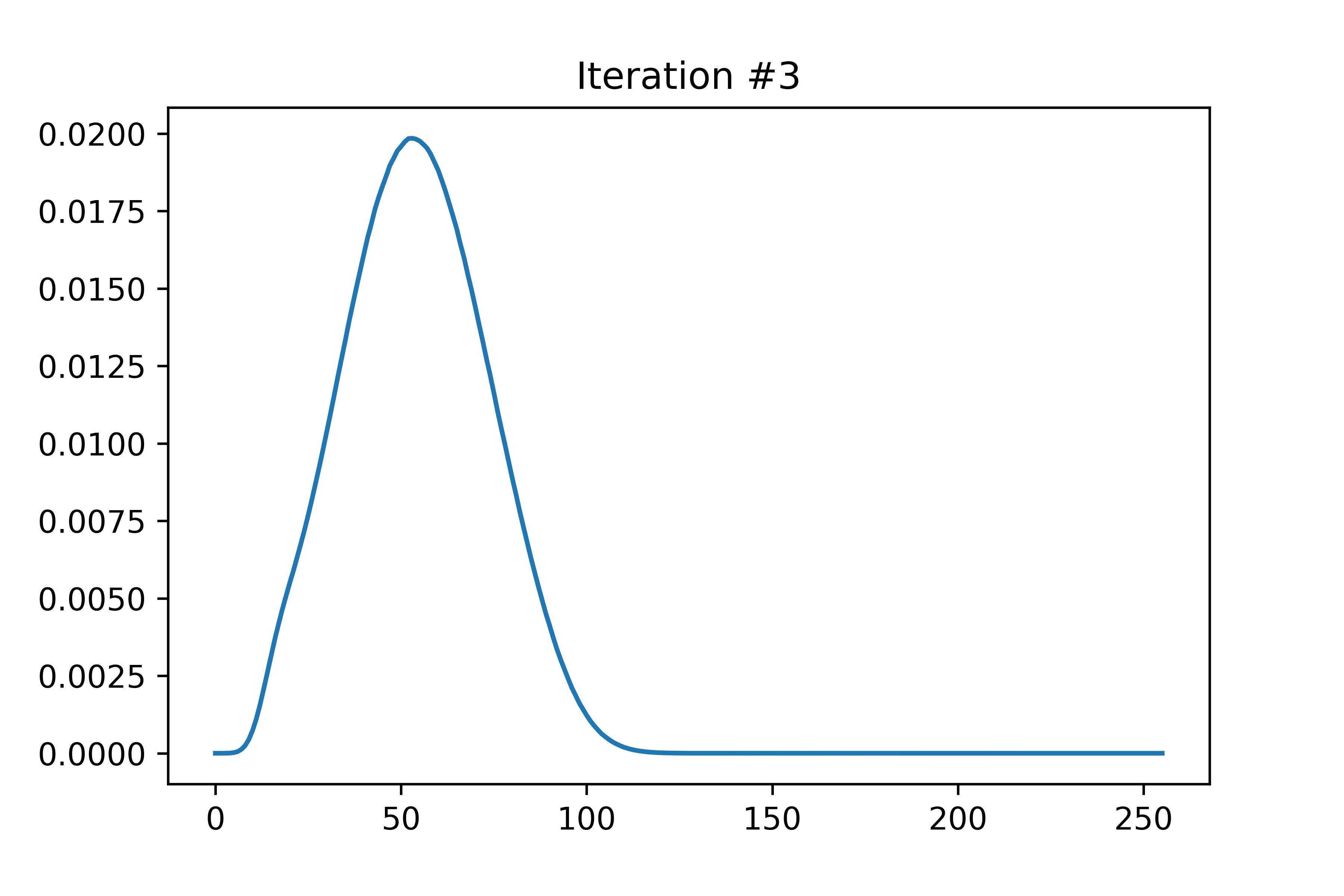
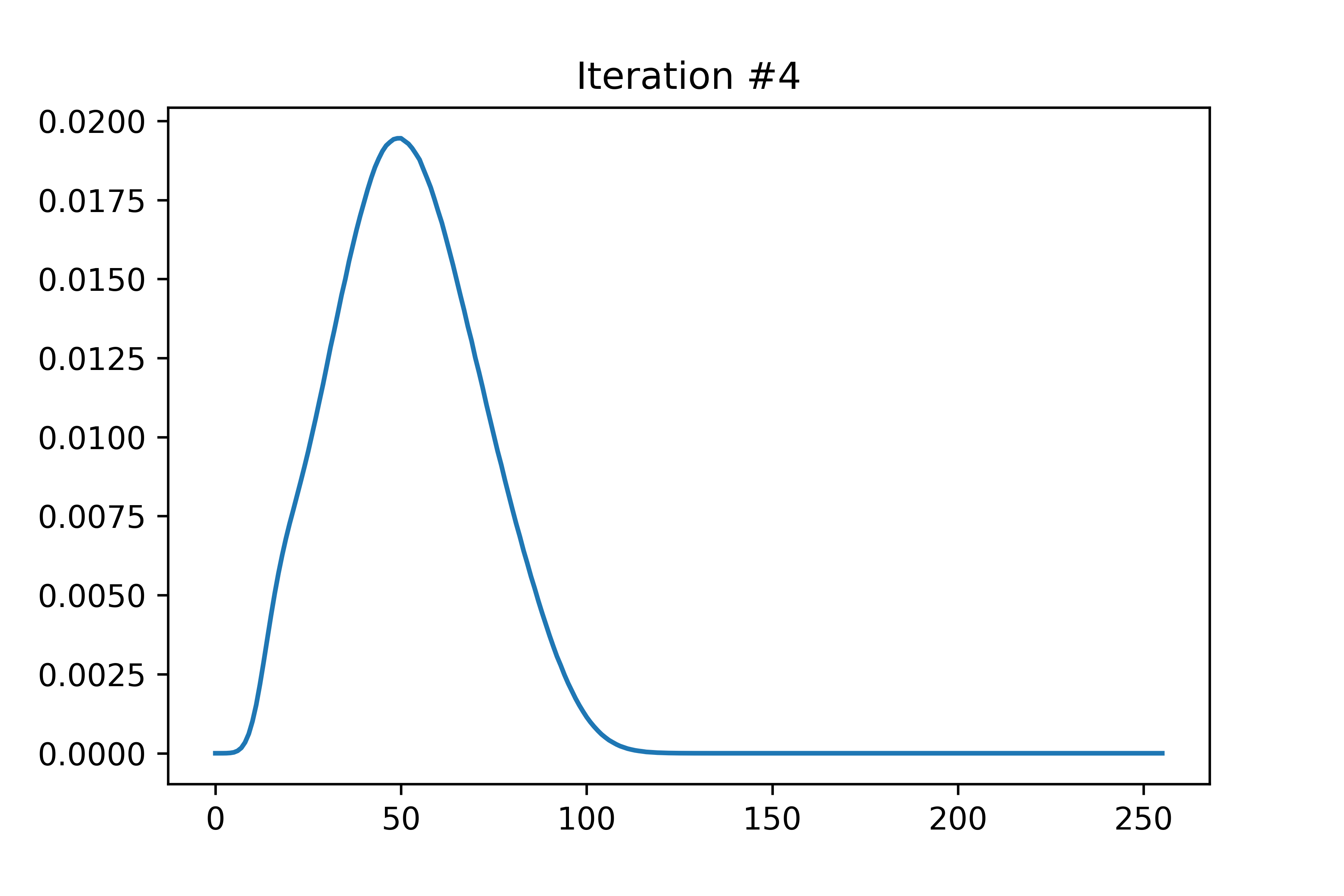
The EDMFT loop starts with a trivial G0_iw, and then iterates. The discrepancy between G['up'] and G['dn'] is not present for the first iteration, then is slightly visible for the second, then is clearly visible for the third and fourth. The Green's functions obtained for the first four iterations is attached below the code. With more iterations, G converges to a spin asymmetric form. The histograms for the four iterations are attached as well.
In the run that produced these plots, S.average_sign was 0.024 for the first iteration, then slightly above 0.20 for the remaining iterations. I used the UNSTABLE branch of CTINT. Each iteration takes about 15-18 minutes, and ran with 800 threads (20 cores) on bnl.
from pytriqs.gf import *
from pytriqs.operators import *
from pytriqs.archive import *
from pytriqs.utility import mpi
from triqs_ctint import Solver
import os
beta = 30.
J = 0.5
U = 6.
mu = U/2.
niter = 4
n_iw = 125
n_tau = 251
n_cycles = 100000
# INITIALIZE
S=Solver(beta = beta,
gf_struct = [('up',[0]), ('dn',[0])],
use_Jperp = True,
use_D = True,
n_tau_dynamical_interactions = n_tau,
n_iw_dynamical_interactions = n_iw)
# G0
for name, g0 in S.G0_iw:
g0 << inverse(iOmega_n + mu - SemiCircular(2))
# DYNAMICAL INTERACTIONS
Qtau = make_gf_from_inverse_fourier(S.Jperp_iw)
# Qtau = S.Jperp_tau.copy()
for i in range(len(Qtau.data)):
Qtau.data[i,0,0] = 0.25
Qiw = make_gf_from_fourier(Qtau)
S.Jperp_iw << -(1./2)*(J**2)*Qiw
S.D0_iw['up','up'] << -(J**2/8.)*Qiw
S.D0_iw['up','dn'] << (J**2/8.)*Qiw
S.D0_iw['dn','up'] << (J**2/8.)*Qiw
S.D0_iw['dn','dn'] << -(J**2/8.)*Qiw
# / INITIALIZE
for iteration in range(niter):
S.solve(h_int = U * n('up',0) * n('dn',0), # Local Hamiltonian
length_cycle = 100,
n_cycles = n_cycles, # Number of QMC cycles
n_warmup_cycles = 5000, # Warmup cycles
n_iw_chi2 = n_iw,
n_tau_chi2 = n_tau,
measure_histogram = True,
measure_chiAB_tau = True,
chi_A_vec = [n('up',0),n('dn',0)],
chi_B_vec = [n('up',0),n('dn',0)],
delta=.2,
)
# SAVE ITERATION
if mpi.is_master_node():
fname="deltatest-i%i.h5"%(iteration)
if os.path.isfile(fname):
os.remove(fname)
with HDFArchive(fname) as A:
A['S']=S
A['h']=S.histogram
A['k']=S.average_k
# / SAVE
# SELF-CONSISTENCY FOR NEXT ITERATION
AvgG = 0.5*(S.G_iw['up']+S.G_iw['dn'])
for name, g0 in S.G0_iw:
g0 << inverse(iOmega_n + mu - AvgG)
Qtau[0,0] << 0.25*(S.chiAB_tau[0,0]+S.chiAB_tau[1,1]-S.chiAB_tau[0,1]-S.chiAB_tau[1,0])
Qiw = make_gf_from_fourier(Qtau)
S.Jperp_iw << -(1./2)*(J**2)*Qiw
S.D0_iw['up','up'] << -(J**2/8.)*Qiw
S.D0_iw['up','dn'] << (J**2/8.)*Qiw
S.D0_iw['dn','up'] << (J**2/8.)*Qiw
S.D0_iw['dn','dn'] << -(J**2/8.)*Qiw
# / SELF